Eublepharis macularius, the leopard gecko, is an animal we have all come to know and love. They are easy to care for and make great pets, but what has become so eye catching about them is the vast diversity of their appearance. Through the many decades of captive breeding of these animals, several traits have randomly mutated and popped out, such as the three strains of albinism (Tremper, Bell, and Rainwater.) Other traits have affected the visual appearance of the eye or the pigmentation patterns on the body. For years now, these traits have been mislabeled as ‘genes’ due to seemingly following simple inheritance patterns (aside from polygenic traits.) The truth is, we have no idea what these traits are. The fact that they are inherited consistently and reproducibly, however, does mean they are part of the genetic material of the animal.
Many new hobbyists, and even veteran breeders in their formative years, likely remember getting told at one point or another to never cross albino strains. However, for people who take the hobby of breeding these remarkable animals seriously, how can one be certain they can trust the breeder from which their animals are coming from to not have crossed strains of albino? For example, there have been instances in the past where breeder ‘X’ buys from breeder ‘Y’, and on instinct, test breeds the animal, only to discover it carries two strains of albino.
In this hobby, that situations like this can go on to ruin projects, result in money down the drain, and if gone unnoticed or untested, could potentially ruin breeder X’s reputation down the line. So how do we, as a hobby, go about ensuring the genetics of our animals, so we can begin to filter the genetic waters that have been murky for some time? The answer is that we physically test the genetics of our animals without breeding.
Disentangling Leo genetics
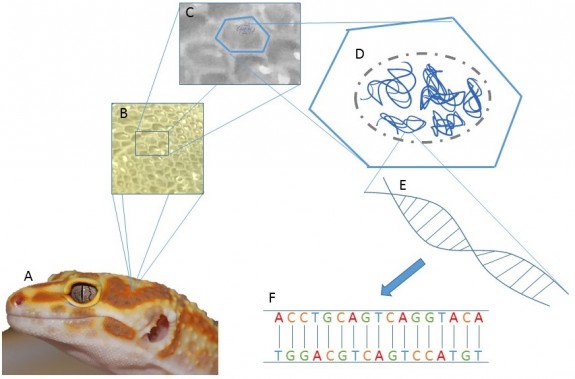
Genotyping is the act of screening an individual animal’s genome and comparing it to a reference to detect differences. Before diving more into this, let me explain some terminology. A genome is all of the genetic material an animal possesses. Every cell of the animal (for the most part) has an entire copy of the animal’s genome, packed into the cell’s nucleus (figure 1A-D.)
Before we will be able to genotype an animal, we first need to know the DNA sequences of the entire genome. Once that information is known, the genomes of multiple animals can be compared, allowing us to identify variations exclusive to each trait. For example, there will be certain variations exclusive to Tremper albino animals compared to wild types, and so on. So how can all of this information be obtained? Through a genome sequencing project.
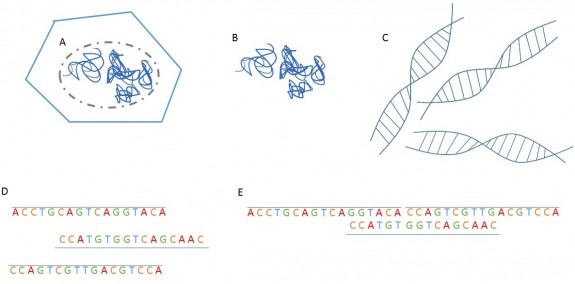
Such a project would involve isolating the DNA from the cells, so that the DNA is the only part of the cells in a sample (figure 1E, 2B.) From there, the DNA can be sequenced and analyzed in a number of fashions. The end result would be the data, being the sequence of the nucleotides in the genome. Nucleotides are the unit of information that makes the genome. There are four nucleotides, which are denoted A, T, C, and G (figure 1F.) To offer an idea of the vastness of a genome, Leopard Gecko genome is estimated to contain over 4,500,000,000 nucleotides1. Once all of the sequencing data is available, it would need to be assembled in a linear fashion (figure 2), much like reading a book. There is currently a draft genome being worked on for leopard geckos, meaning that the sequencing is not yet finished, and the next step would be assembly of the sequences.
Once the reference genome is assembled, the genomes of animals with pure traits (i.e. pure Blizzard, with no heterozygosity) will be compared to the reference to identify the exclusive differences. The identification of such differences is not guaranteed, or even highly likely, and may often require samples to be run multiple times before finding the differences sought after. This task is not one that will be easy, or cheap, but at least needs to be attempted to advance our beloved hobby. Once all of these differences are identified, animals will be able to be genotyped. However, keep in mind, that the process will be a long and winding road, filled with hurdles, troubleshooting, insignificant results, and so on.
What would this mean for the breeding hobby?
Once animals can be genotyped, it will truly revolutionize the breeding hobby. There will be no more unnecessary distrust about unknown heterozygosity, or crossed albinos. Breeders would no longer need to feel compelled to test breed, therefore minimizing the unnecessary offspring produced from such breeding. Another major change would be that animals would no longer need to go on labeled as ‘possible het’, but instead, could be tested and the buyer could know for sure, one way or another.
Perhaps the two most important effects this could have on the hobby, would be to clean up many lines to clear the murky genetic waters, and to potentially identify the actual genetic components responsible for Enigma Syndrome (ES.) Being able to test animals for multiple albinisms without breeding them and creating countless babies as a byproduct will be infinitely priceless. However, even more priceless is the fact that this would make reestablishing pure lines much easier, to ensure they will be around for the generations to come.
If the project is successful at identifying the genetic basis of ES, there are a few possible scenarios: 1) the genetic material responsible for ES is actually part of the enigma trait, 2) the previous theory of ES being closely associated through linkage disequilibrium could be proven (see Ben Bargen’s Gecko Time article on ES) If the latter proves true, the possibility of eliminating ES from lines will become very real. Additionally, and perhaps most importantly, if the molecular basis of ES is known, it can lead to a collaboration with veterinary and pharmaceutical staff to develop potential treatments for animals who are affected by ES.
What is the status of such a project?
As stated earlier, there is a draft genome being worked on (not through myself.) This project is still in the hypothetical state, meaning there is a lot of planning, discussing, troubleshooting, and trying to determine the best course of action to achieve the goals. Once more planning is done and preliminary results become available, there will be a follow up article, and a blog will be put together so the general public can follow along with progress as it is made. In the meantime, a Facebook group is being put together now, called Leopard Gecko Genetics, solely for the discussion of these topics.


This is a topic I’ve been somewhat interested in myself. The school I attend has some dna sequencing equipment, and I’ve got two unrelated male leopard geckos. I don’t know if I could afford to buy instrument time on my own, but I am interested in getting in touch with a group and seeing if there is any way I could help out. The Facebook link doesn’t seem to be working, though.
Hi JP,
I’m not sure why the link isn’t working, but here is the direct link to the FB group I started; https://www.facebook.com/groups/345210672310425/
At this point, given that there is a draft genome being worked on through an external project, it will be a waiting game for that assembly to be drafted to be able to compare the various traits to. I am meeting with a technician and representative of a sequencing company next Wednesday to discuss the project and go over the aims, as well as the various sequencing methods available through the company.
Once I have more of the project planned out, methodologies chosen and started, I will do a follow up article in more detail for those interested.
Best,
Dillon
Am wondering if the draft genome will be based upon genetic sampling of wild leopard gecko populations for the baseline?
Regards,
Jeff
Hi Jeff,
To the best of my knowledge, the draft genome is being done with samples from bloodlines directly derived from wild caught animals, but inbred for a few generations to reduce some heterogeneity of the gene pool.